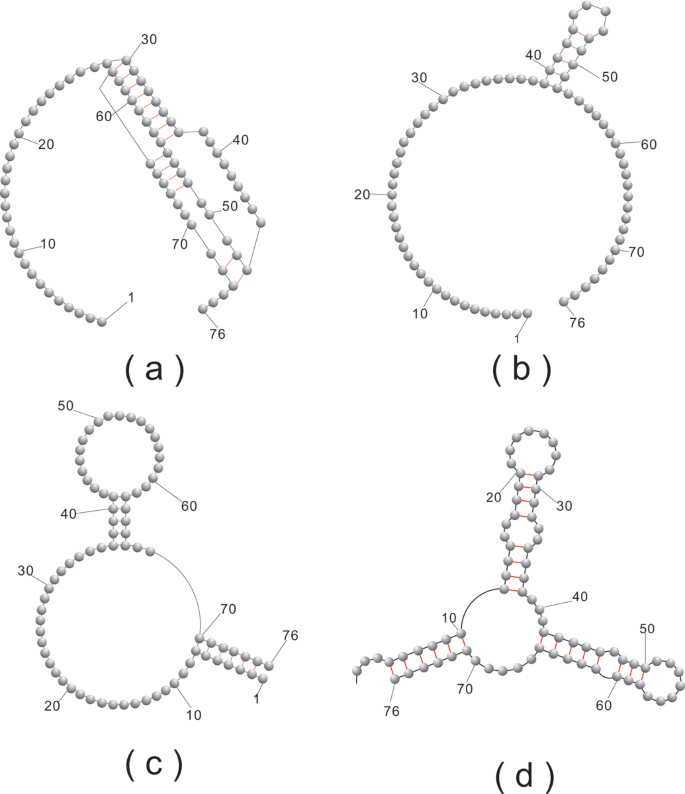
Mapping in Solution Shows the Peach Latent Mosaic Viroid To Possess a New Pseudoknot in a Complex, Branched Secondary Structure | Journal of Virology

Small molecule targeting of biologically relevant RNA tertiary and quaternary structures - ScienceDirect
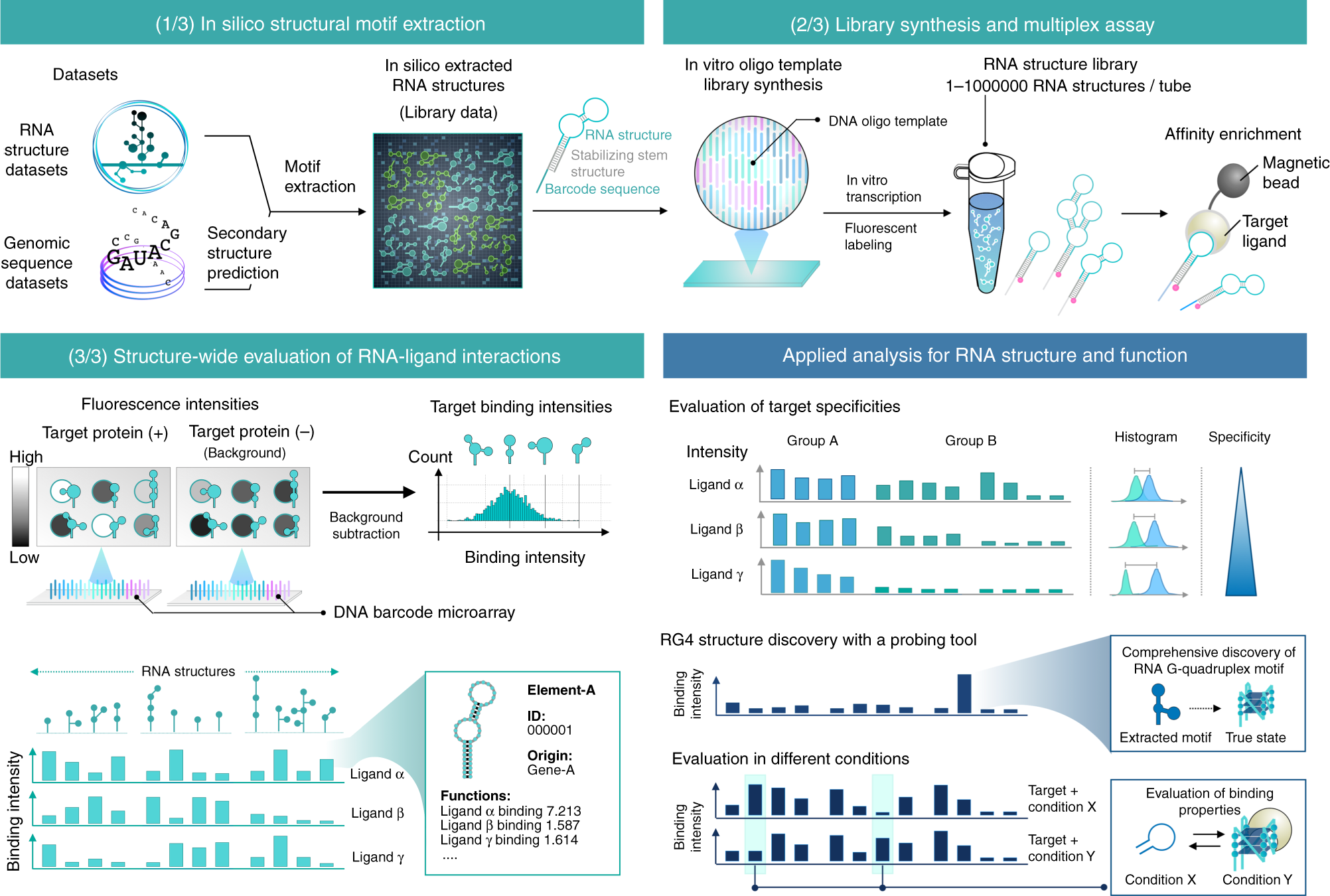
RNA structure-wide discovery of functional interactions with multiplexed RNA motif library | Nature Communications

An RNA secondary structure illustrating the types of features included... | Download Scientific Diagram

An RNA secondary structure with examples of the five kinds of loops:... | Download Scientific Diagram
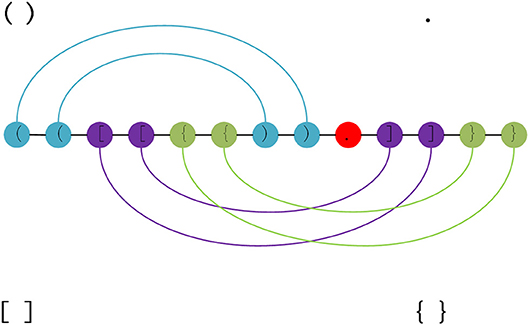
Frontiers | DMfold: A Novel Method to Predict RNA Secondary Structure With Pseudoknots Based on Deep Learning and Improved Base Pair Maximization Principle

Structures of artificially designed discrete RNA nanoarchitectures at near-atomic resolution | Science Advances

Structural dynamics of the SARS-CoV-2 frameshift-stimulatory pseudoknot reveal topologically distinct conformers | bioRxiv
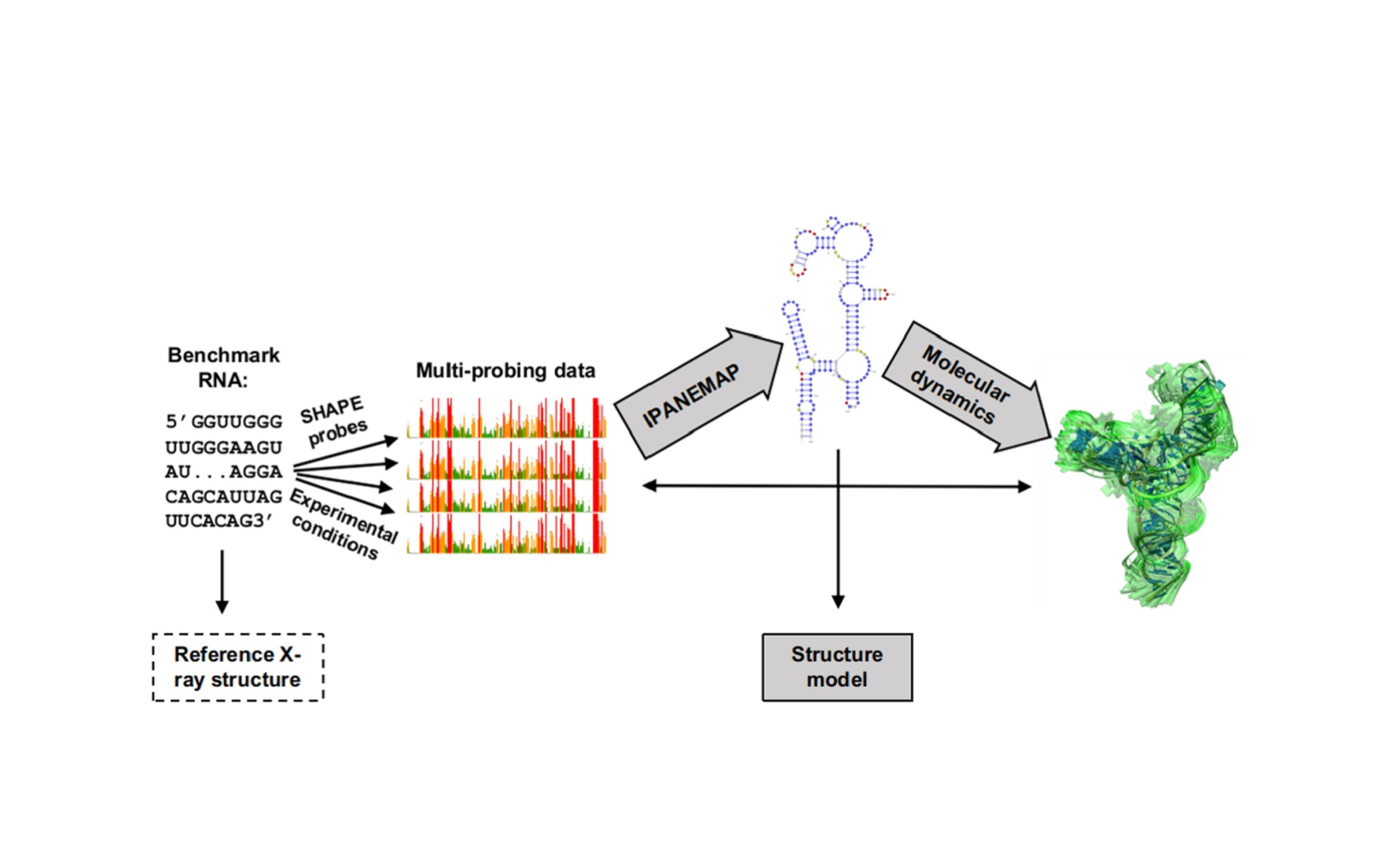
ncRNA | Free Full-Text | Progress toward SHAPE Constrained Computational Prediction of Tertiary Interactions in RNA Structure
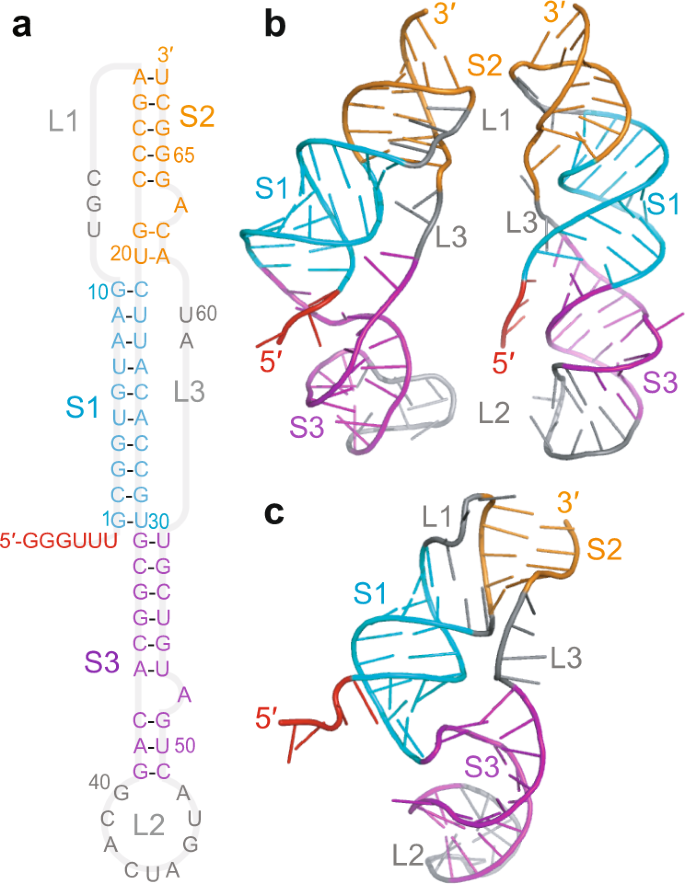
Structural dynamics of single SARS-CoV-2 pseudoknot molecules reveal topologically distinct conformers | Nature Communications
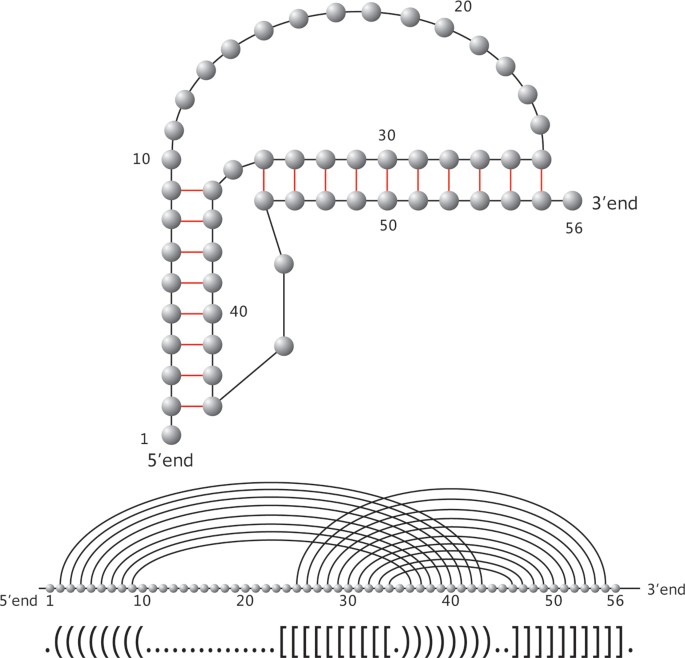
Prediction of RNA Pseudoknots Using Heuristic Modeling with Mapping and Sequential Folding | PLOS ONE
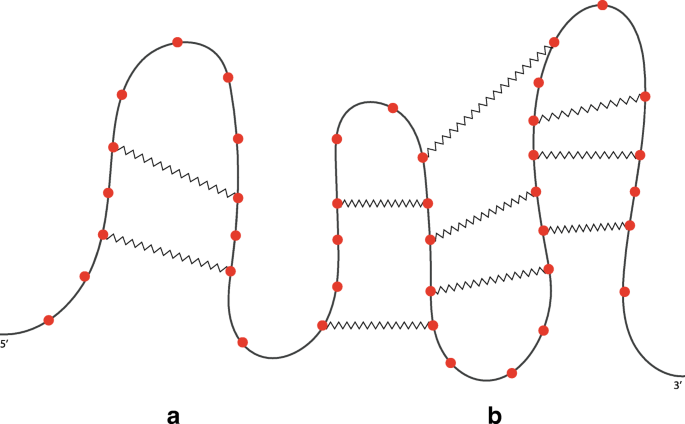
RNA structure types. (A) cartoon schematic of RNA structure types. (B)... | Download Scientific Diagram

Mechanical unfolding kinetics of the SRV-1 gag-pro mRNA pseudoknot: possible implications for −1 ribosomal frameshifting stimulation | Scientific Reports

Multistrand RNA Secondary Structure Prediction and Nanostructure Design Including Pseudoknots | ACS Nano
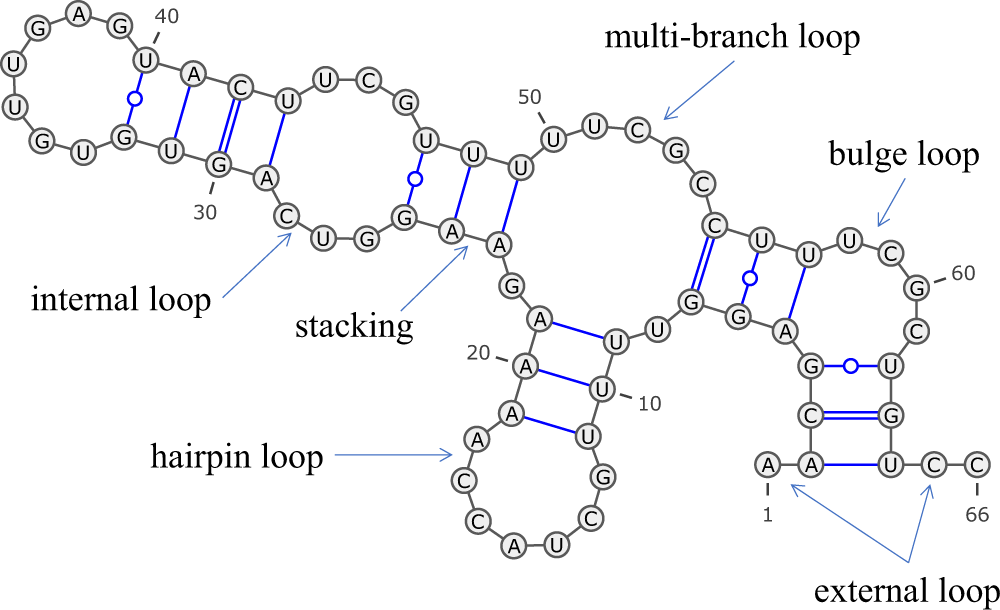
RNA secondary structure prediction using deep learning with thermodynamic integration | Nature Communications
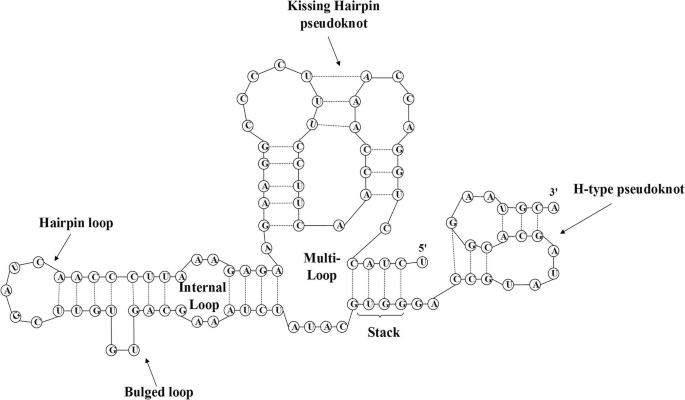
An efficient simulated annealing algorithm for the RNA secondary structure prediction with Pseudoknots | BMC Genomics | Full Text

RNA structure types. (A) cartoon schematic of RNA structure types. (B)... | Download Scientific Diagram